Dominique LE COQ | Researcher | PhD | French National Institute for Agriculture, Food, and Environment (INRAE), Paris | INRAE | Research profile

Structure–Kinetic Relationship for Drug Design Revealed by a PLS Model with Retrosynthesis-Based Pre-Trained Molecular Representation and Molecular Dynamics Simulation | ACS Omega

Surfactin, a quorum sensing signal molecule, globally affects the carbon metabolism in Bacillus amyloliquefaciens - ScienceDirect

Surfactin, a quorum sensing signal molecule, globally affects the carbon metabolism in Bacillus amyloliquefaciens - ScienceDirect

Inhibition of CPAP–tubulin interaction prevents proliferation of centrosome‐amplified cancer cells | The EMBO Journal
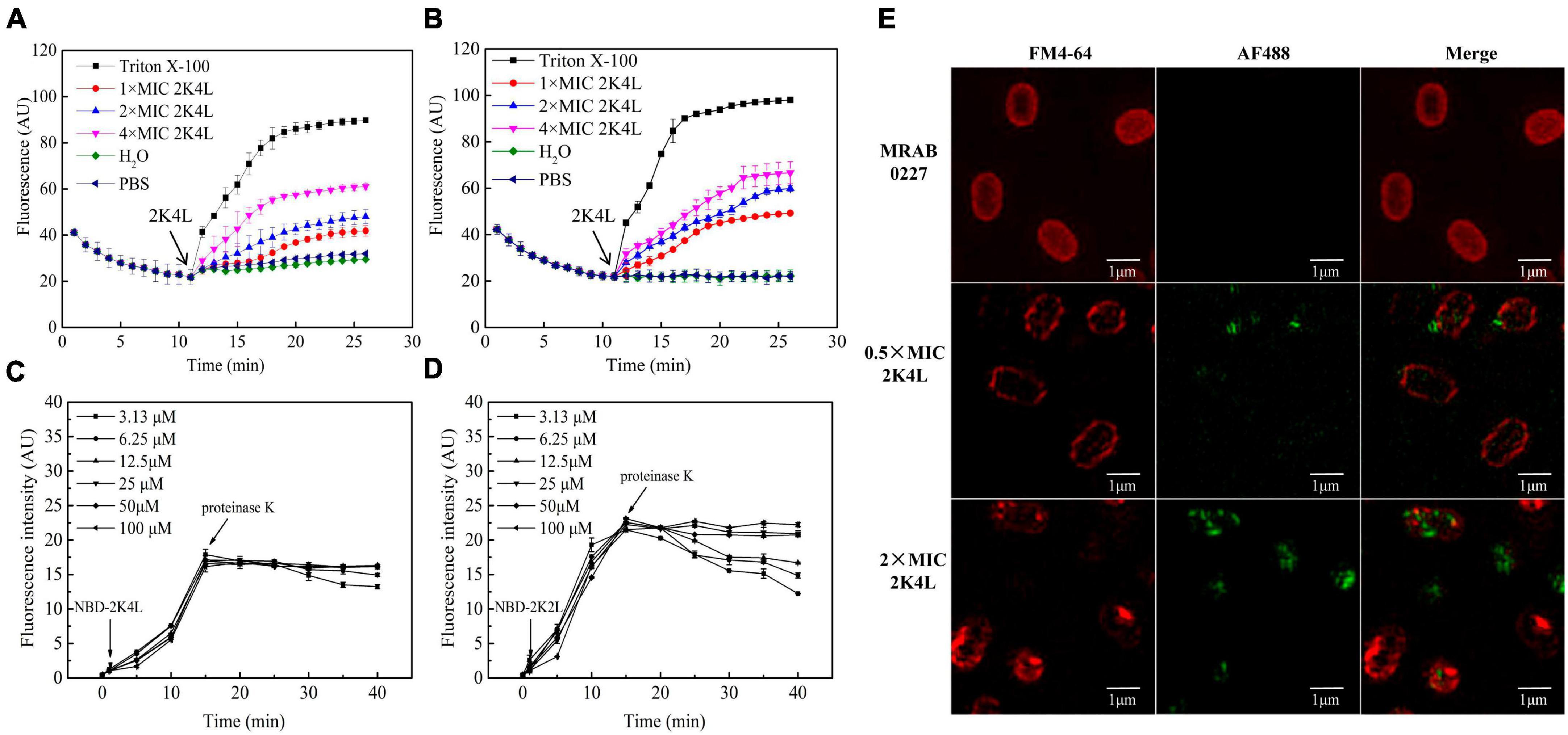
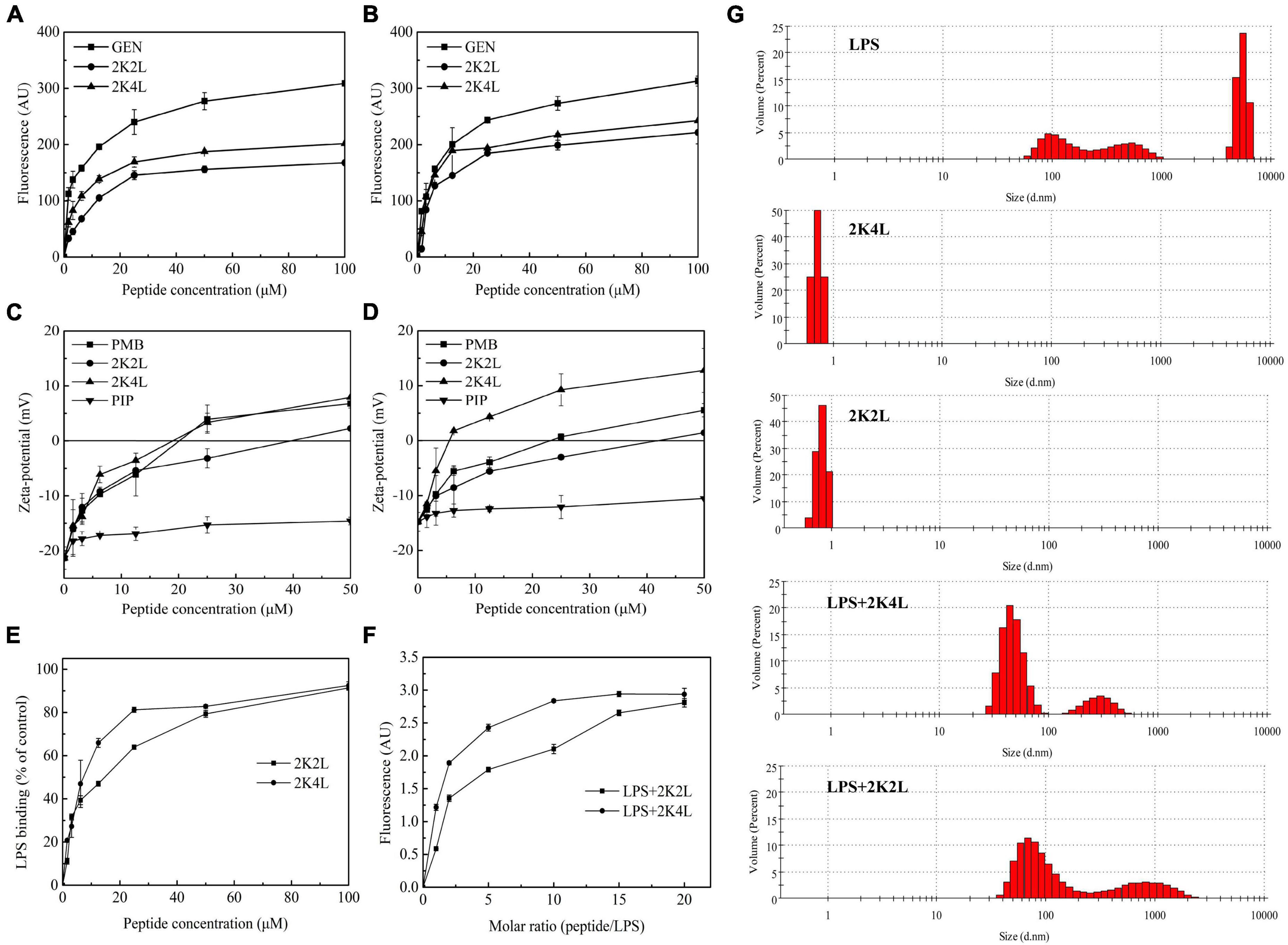
Frontiers | Antimicrobial peptide 2K4L disrupts the membrane of multidrug-resistant Acinetobacter baumannii and protects mice against sepsis

PDF) Hsp104 N‐terminal Domain Interaction with Substrates Plays a Regulatory Role in Protein Disaggregation

Phosphorylation of a Borealin Dimerization Domain Is Required for Proper Chromosome Segregation | Biochemistry

Acidic Residues Control the Dimerization of the N-terminal Domain of Black Widow Spiders' Major Ampullate Spidroin 1 | Scientific Reports

Catalysts | Free Full-Text | Flexibility and Function of Distal Substrate-Binding Tryptophans in the Blue Mussel β-Mannanase MeMan5A and Their Role in Hydrolysis and Transglycosylation
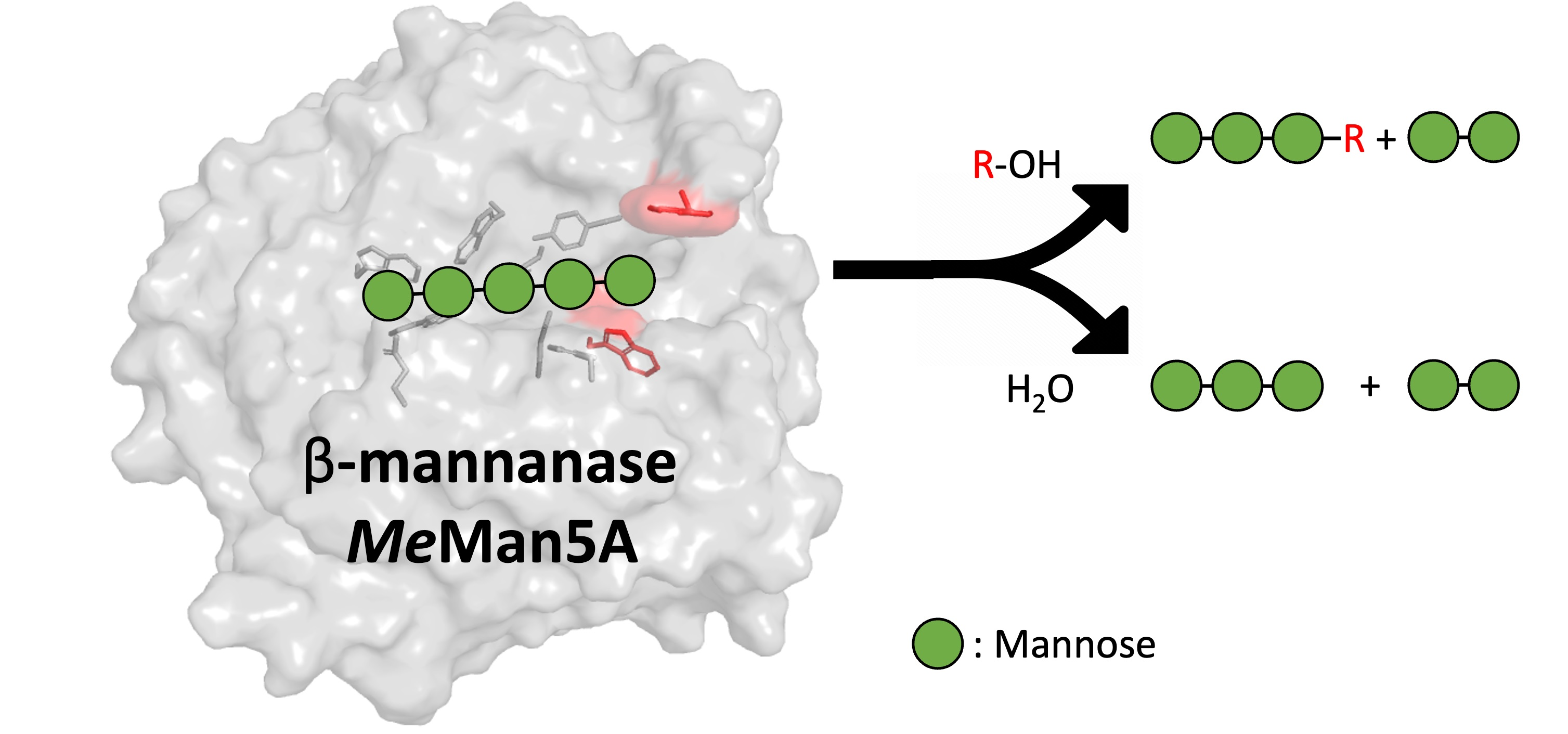
Catalysts | Free Full-Text | Flexibility and Function of Distal Substrate-Binding Tryptophans in the Blue Mussel β-Mannanase MeMan5A and Their Role in Hydrolysis and Transglycosylation
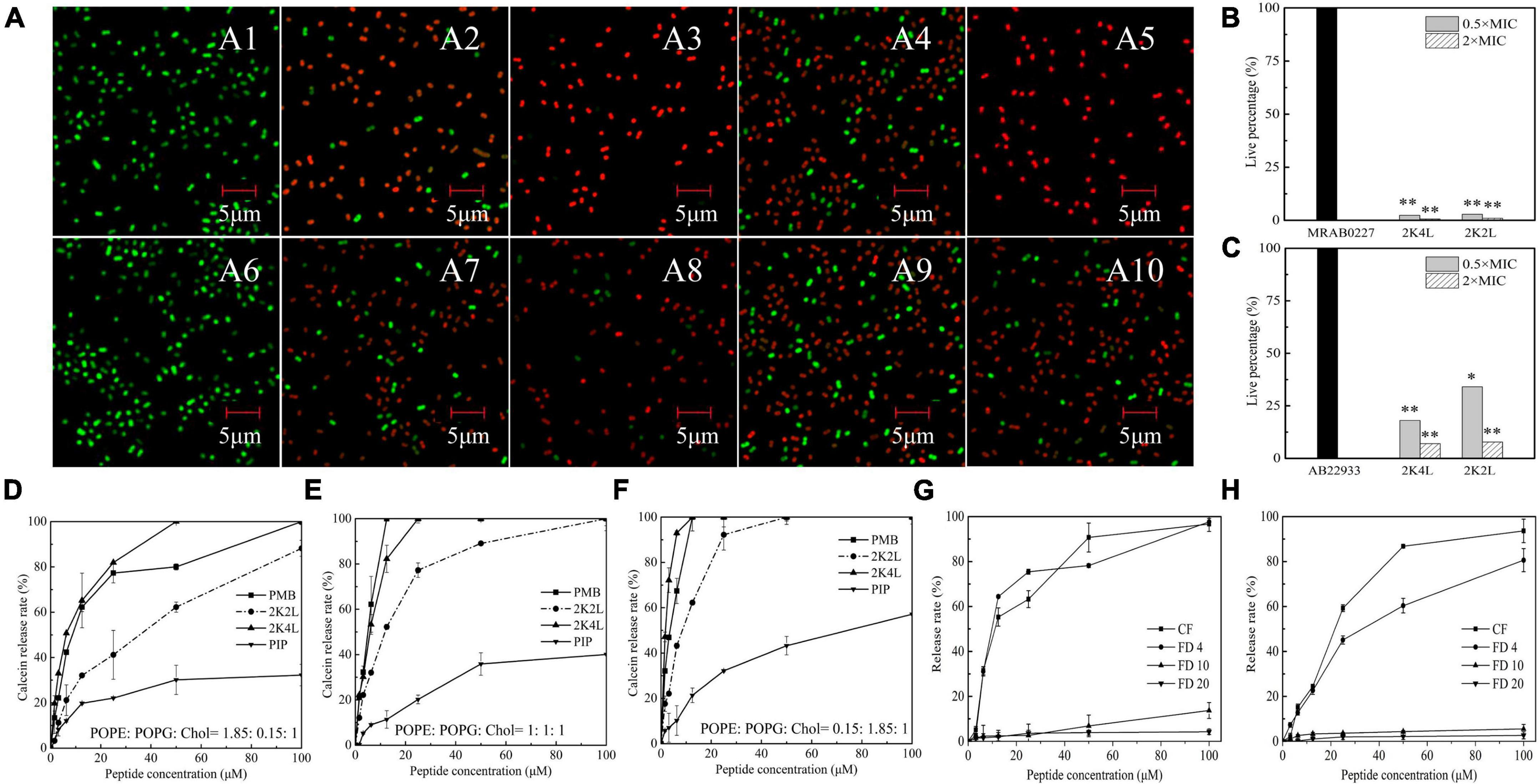
Frontiers | Antimicrobial peptide 2K4L disrupts the membrane of multidrug-resistant Acinetobacter baumannii and protects mice against sepsis
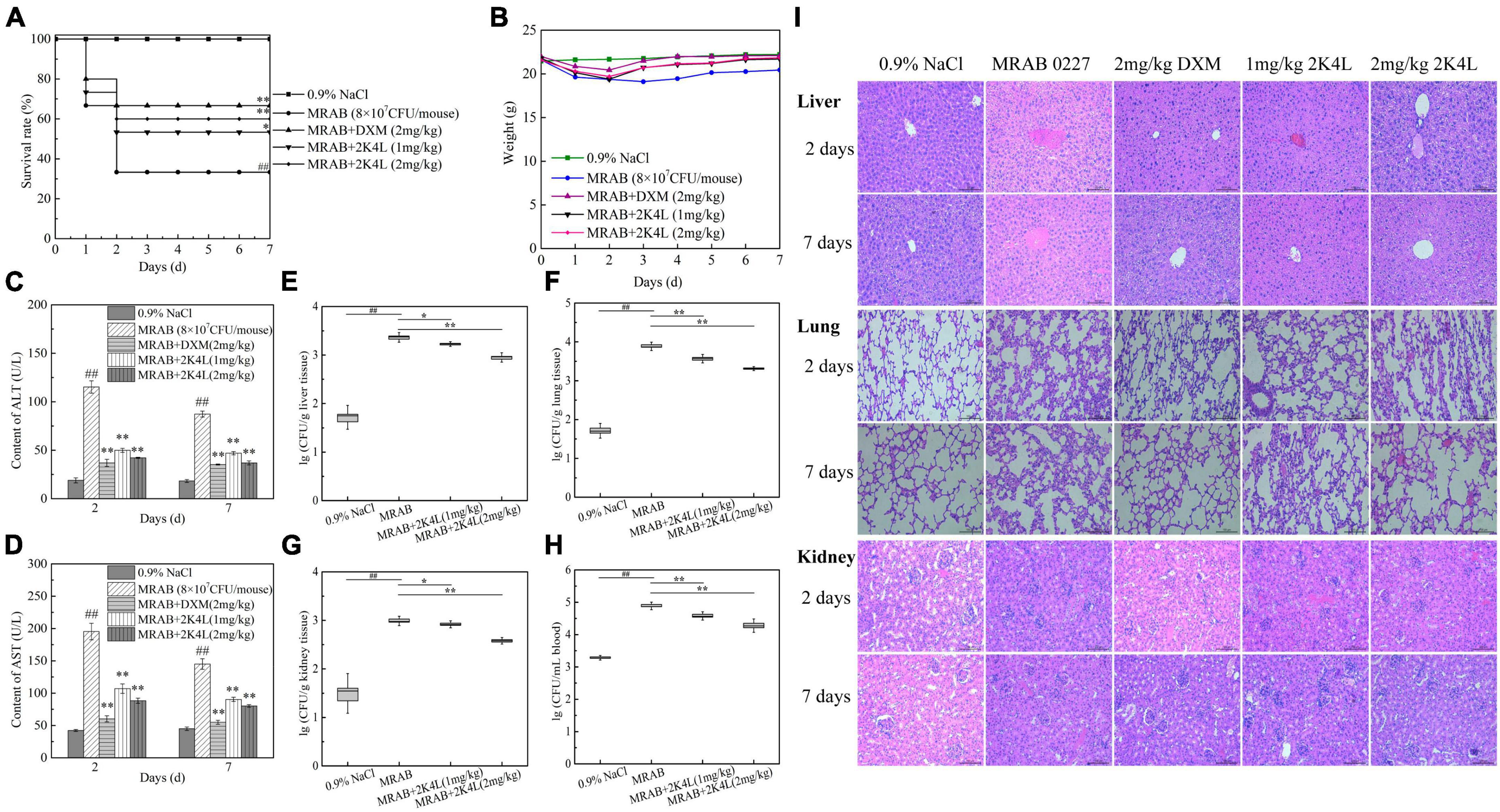
Frontiers | Antimicrobial peptide 2K4L disrupts the membrane of multidrug-resistant Acinetobacter baumannii and protects mice against sepsis

Structure–Kinetic Relationship for Drug Design Revealed by a PLS Model with Retrosynthesis-Based Pre-Trained Molecular Representation and Molecular Dynamics Simulation | ACS Omega